R
install.packages('tidyverse')
install.packages('patchwork')
install.packages('ggrepel')
install.packages("pheatmap")
install.packages("tidyheatmaps")
install.packages("BiocManager")All bioinformatics projects requires setting up the required software. Let’s go through the setup for this RNA-seq workshop here.
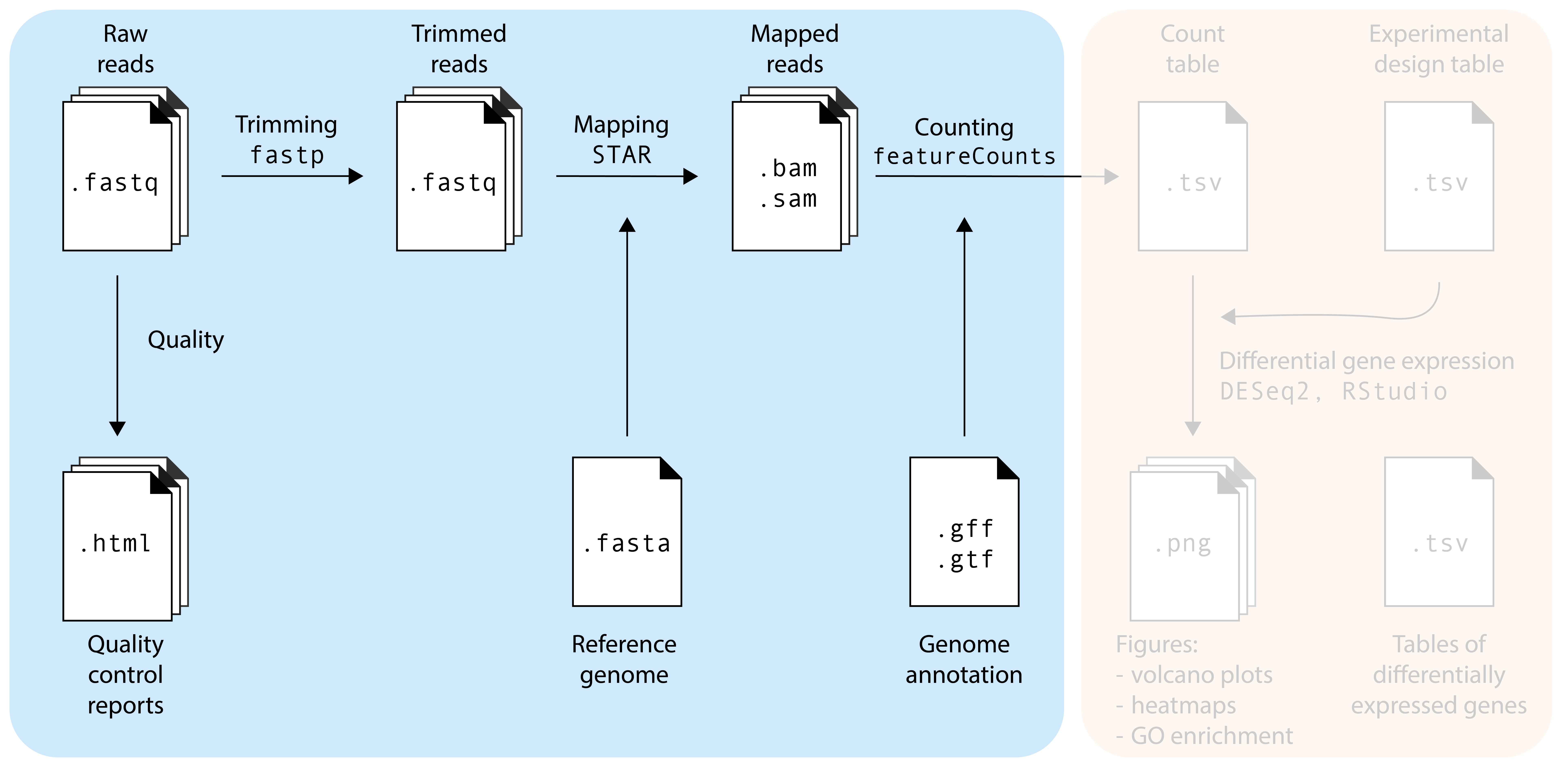
The first part is the ‘hardcore’ bioinformatics part of this workshop. These steps are nearly always performed using command-line programs, which can be accessed via the Terminal (macOS, Linux) or WSL (Windows). For this workshop, we prepared a Google Colab environment where we will run the commands. Access to the Google Colab environment requires a Google account.
In a real experiment, you will often have so many samples and reads that you will need access to a compute cluster with more compute power (CPUs), working memory (RAM), and storage space. At the University of Amsterdam, we use Crunchomics. Crunchomics comes with many popular bioinformatics pre-installed, additional programs can be installed using conda.
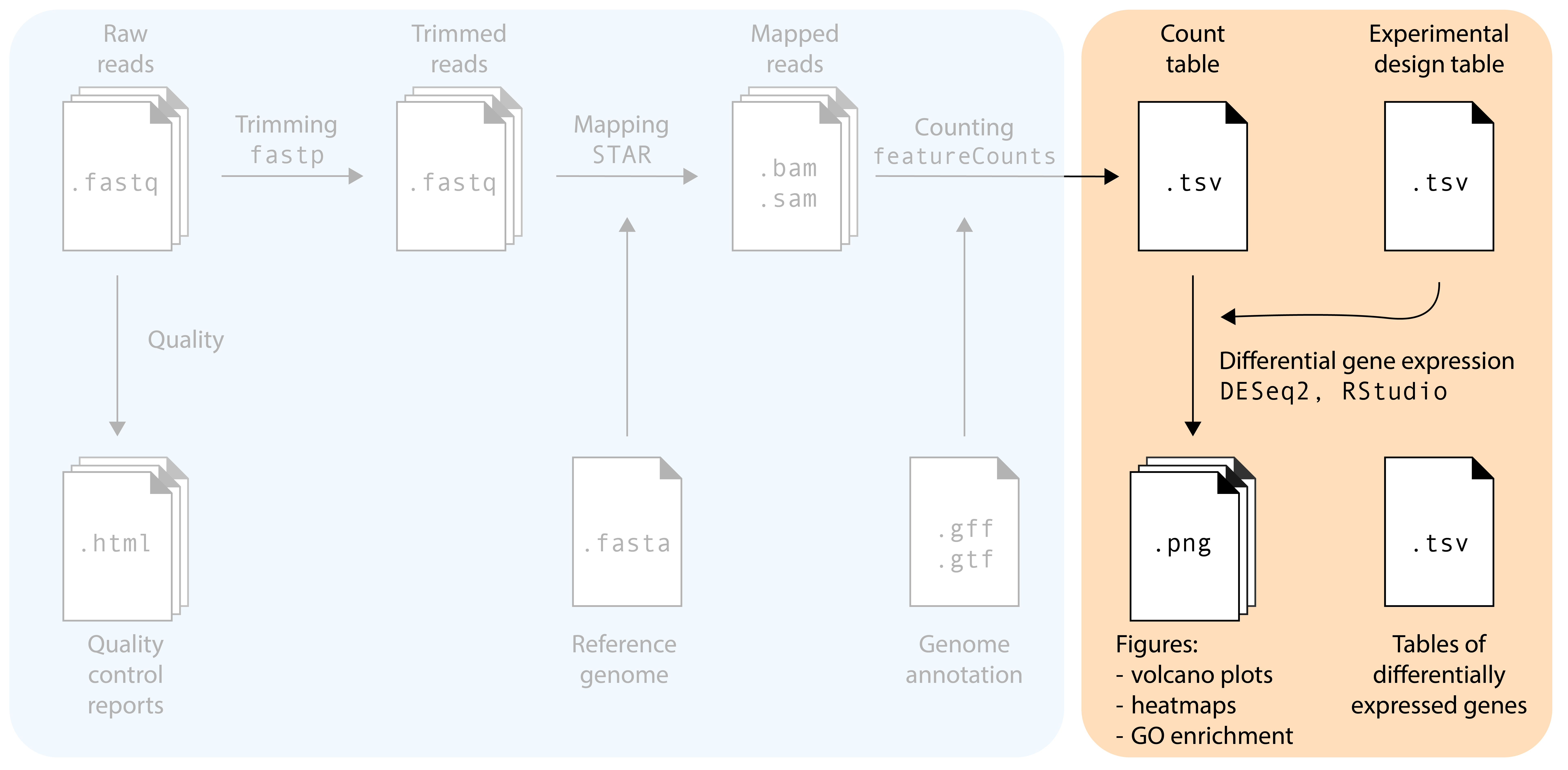
For the second of the workshop, we will use R. Please make sure you have R and RStudio installed on your computer. In addition, you will need to install the following packages, via two different routes:
Install the following packages via install.packages()
R
install.packages('tidyverse')
install.packages('patchwork')
install.packages('ggrepel')
install.packages("pheatmap")
install.packages("tidyheatmaps")
install.packages("BiocManager")Install the following packages via Bioconductor
R
BiocManager::install("apeglm")
BiocManager::install("enrichplot")
BiocManager::install("clusterProfiler")
BiocManager::install("org.At.tair.db")
BiocManager::install("DESeq2")
BiocManager::install("EnhancedVolcano")Test whether DESeq2 was installed properly by loading it:
R
library(DESeq2)During the workshop, we will not have time to install the packages, or to troubleshoot installation problems. Please try to install the packages before the workshop. You can always contact the workshop organizers if you run into problems.
The reads and reference that will be used in this workshop can be found in the following Github repository, and will be downloaded into the Google Colab automatically.
The count table and experimental design file used in the DESeq2 part of this workshop can be found here.